DNA methylation at CpG dinucleotides, the most common DNA modification in eukaryotes, has been associated with various phenomena such as alterations in chromatin structure, genomic imprinting, transposon and chromosome X inactivation, differentiation, and cancer. Effects of DNA methylation are mediated through proteins that bind to symmetrically methylated CpGs. Such proteins contain a specific methyl-CpG-binding domain (MBD). The Methyl-CpG-binding domain (MBD) binds to DNA that contains one or more symmetrically methylated CpGs, while it has a negligible non-specific affinity for unmethylated DNA.
Methylated sequence capturing using a MBD based enrichment followed by next-generation sequencing thus provides a great combination of sensitivity and cost-efficiency for genome-wide DNA-methylation profiling.
Advantages of MBD Sequencing
- True genome-wide
- Approximate basepair resolution
- 100 times more sensitive than MeDIP
- >95% of genome-wide bisulfite sequencing data is also found with MBD sequencing
- Low DNA input (500ng genomic DNA)
- Applicable to all mammalian samples and tissue types
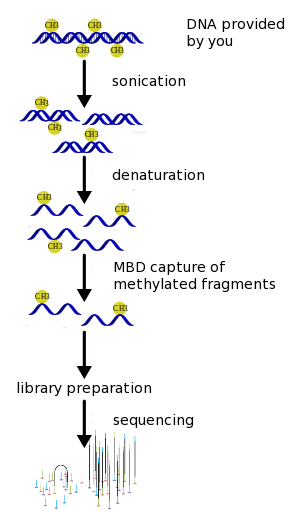
MBD Sequencing Workflow
As for all our sequencing services, we start with performing a first quality control step in which we measure the quantity and quality of the DNA provided to us by our customers. Only when the provided genomic DNA is of sufficient quality and quantity will we proceed with the next steps. In case the DNA provided to us is of insufficient quality and/or quantity, we will contact the customer and discuss how to proceed.
Then, we perform a random fragmentation of the genomic DNA samples followed by size selection of the resulting fragments. This is followed by a second quality control step in which the resulting DNA concentration is determined and the DNA is checked for degradation.
In a third step, the resulting DNA fragments will then be separated by performing MBD-capture during which all fragments containing methylation are withheld, while fragments that do not contain methylation are discarded.
Finally, the library preparation and sequencing are then performed using one of our Illumina sequencers. The technical quality of the sequencing run is monitored in real time.
Sequencing data can be transferred to you via the Illumina BaseSpace platform or our own server (FTP download).
If you have more questions, please feel free to contact us.